Lipid metabolism during growth and stress
|
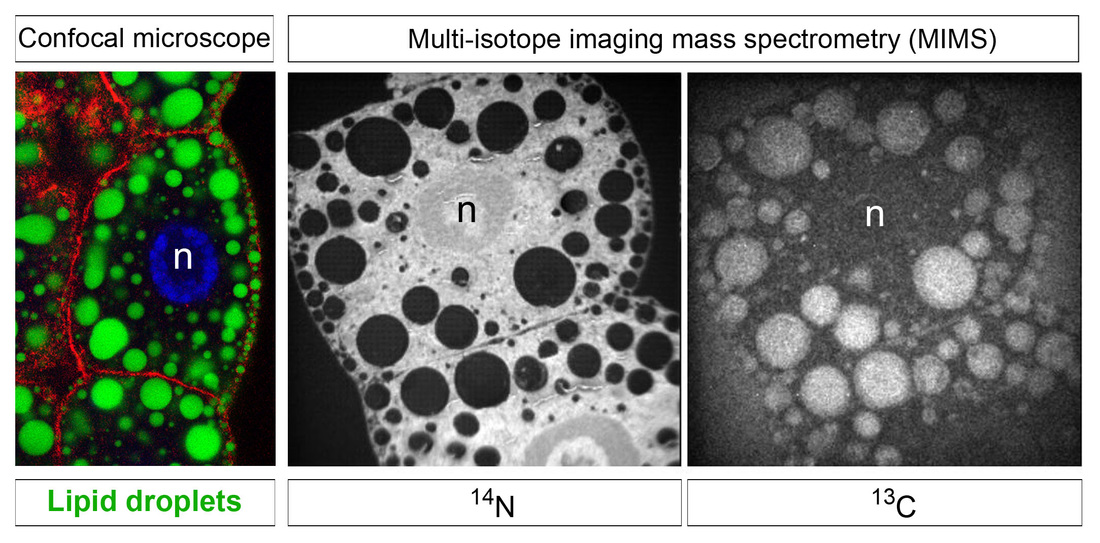
Lipids play numerous different roles in health and disease. Some types of lipids function as metabolic fuels for energy homeostasis, others act as components of cell membranes and some serve as signalling molecules. Neutral lipids relevant for energy homeostasis, such as triglycerides and cholestryl esters, are stored inside cells within lipid droplets. These lipid droplets are important organelles consisting of a neutral lipid core surrounded by a phospholipid monolayer containing lipases and many other proteins regulating fat metabolism. Current work in the lab explores the physiological roles of lipid droplets in a number of different biological contexts and disease models.
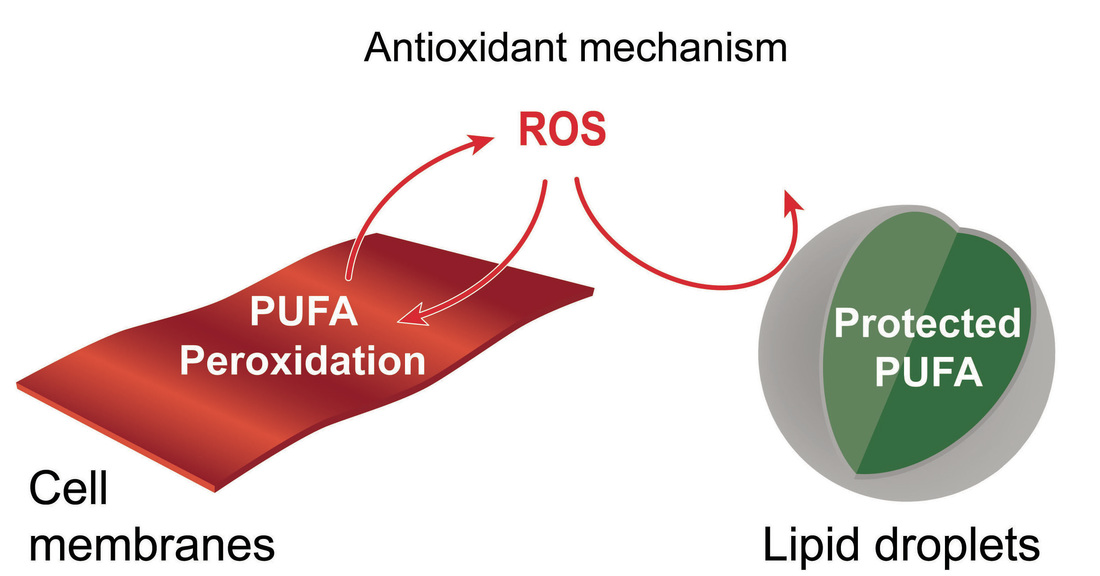
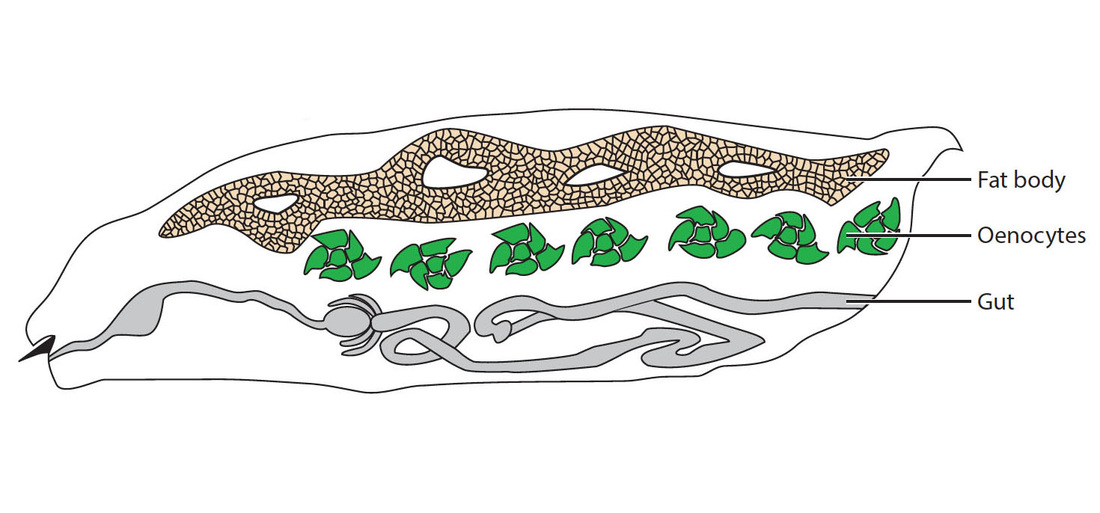
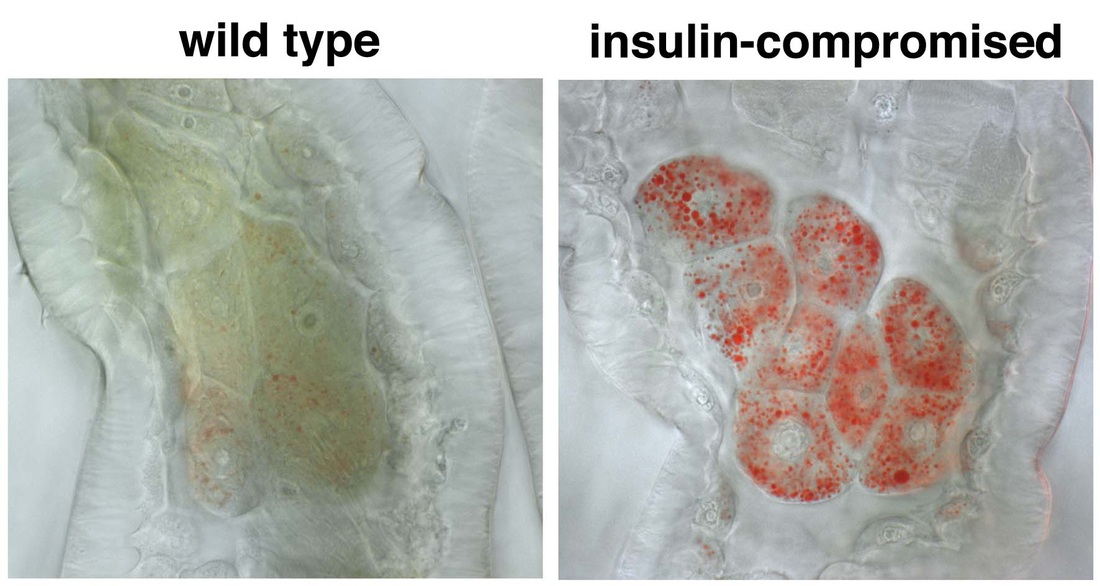
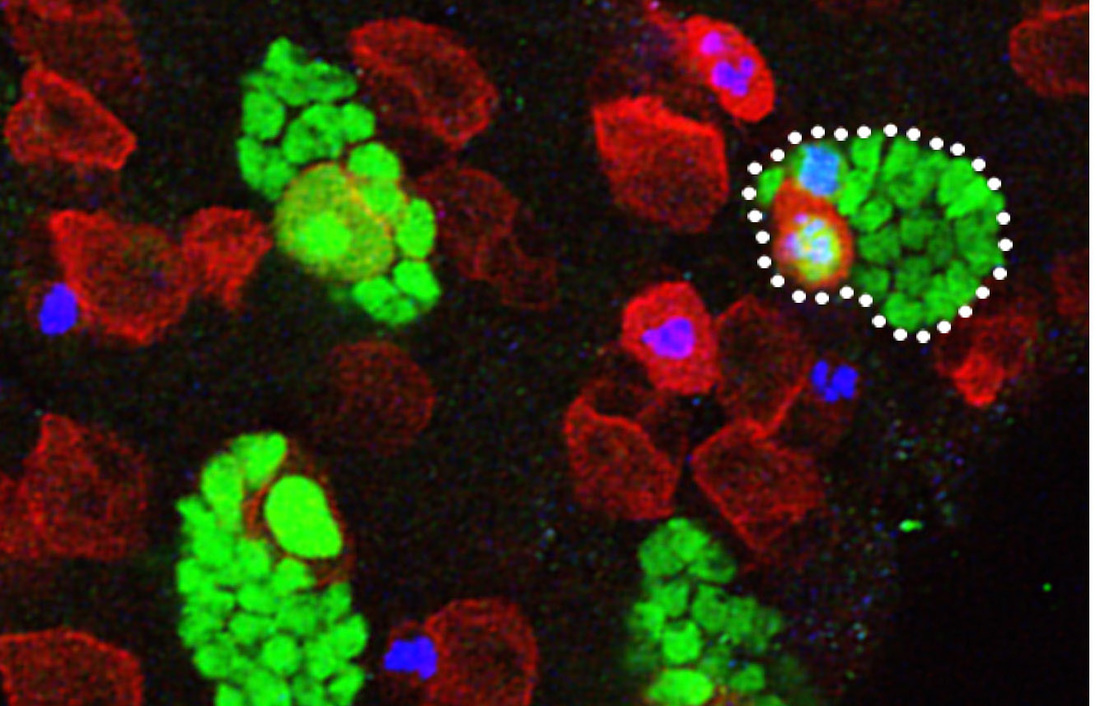
Research in the lab has identified an unexpected role for lipid droplets as an antioxidant organelle. This research began with the observation that lipid droplets in the developing Drosophila CNS are induced by reactive oxygen species (ROS) generated by environmental exposure to hypoxia and other pro-oxidants. Lipid droplets form in the glia of the stem cell (neuroblast) niche during oxidative stress and function to limit ROS levels and to inhibit the peroxidation of polyunsaturated fatty acids (PUFAs). These droplets protect glia and also neuroblasts from peroxidation chain reactions that can damage many types of macromolecules. The underlying antioxidant mechanism involves diverting PUFAs, including diet-derived linoleic acid, away from membranes to the core of lipid droplets, where they are less vulnerable to peroxidation. We are now testing whether the antioxidant role for lipid droplets is relevant in other Drosophila and mammalian contexts. Oenocytes are mysterious insect cells that were first observed in the 1850s but whose functions remained elusive until the 2000s. We discovered that Drosophila oenocytes express genes indicative of highly specialized role(s) in fatty acid metabolism and that they are essential for the growth of the larva. We found that they accumulate lipid droplets during starvation by a mechanism that requires fat mobilization from the major insect adipose store, the fat body. Hence, at least in terms of nutritional regulation, there are interesting similarities between the mammalian adipose-liver and the insect fat body-oenocyte axes. The physiological functions of oenocytes appear to be linked to their ability to synthesize very long chain fatty acids (VLCFAs). Other labs have shown that, in larvae, oenocyte VLCFAs are involved in tracheal waterproofing and that, in adults, they are metabolized into pheromones. Our current work in this area focuses on identifying the mechanisms by which dietary nutrients regulate oenocytes. |
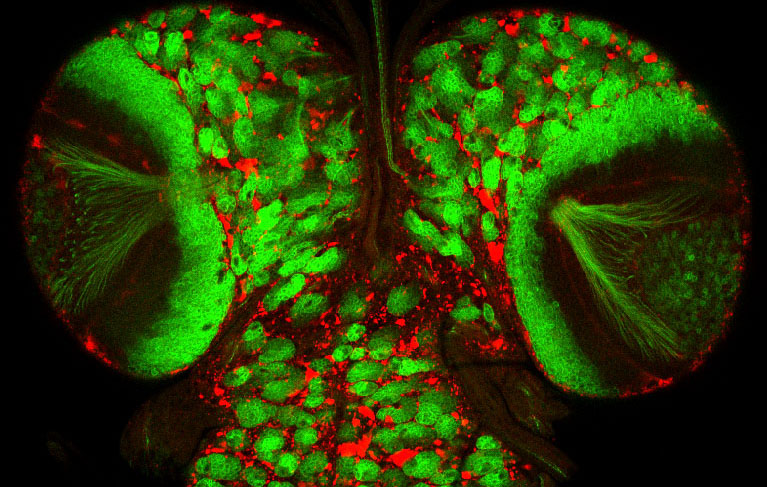
Larval oenocytes (red) are closely associated with the PNS (green)
|
Relevant publications:
Developmental diet regulates Drosophila lifespan via lipid autotoxins
Nature Communications 8, Article number: 1384
M. Irina Stefana, Paul C. Driscoll, Fumiaki Obata, Ana Raquel Pengelly, Clare L. Newell, James I. MacRae, Alex P. Gould (2017)
Lipid droplet functions beyond energy storage
Biochimica et Biophysica Acta (BBA) - 1862(10 Pt B),1260-1272
Michael A. Welte, Alex P. Gould (2017)
Drosophila Spidey/Kar Regulates Oenocyte Growth via PI3-Kinase Signaling
PLoS Genetics 12(8): e1006154
Einat Cinnamon, Rami Makki, Annick Sawala, Leah P. Wickenberg, Gary J. Blomquist, Claus Tittiger, Ze'ev Paroush, Alex P. Gould (2016)
Antioxidant Role for Lipid Droplets in a Stem Cell Niche of Drosophila
Cell 163, 340-353
Bailey AP , Koster G, Guillermier C, Hirst EMA, MacRae JI, Lechene CP, Postle AD, Gould AP (2015)
The development and functions of oenocytes
Annu Rev Entomol 59, 405-25
Makki R, Cinnamon E and Gould AP (2014)
Specialized hepatocyte-like cells regulate Drosophila lipid metabolism
Nature 445, 275-280
Gutierrez, E., Wiggins, D., Fielding, B. and Gould, A.P. (2007)
Developmental diet regulates Drosophila lifespan via lipid autotoxins
Nature Communications 8, Article number: 1384
M. Irina Stefana, Paul C. Driscoll, Fumiaki Obata, Ana Raquel Pengelly, Clare L. Newell, James I. MacRae, Alex P. Gould (2017)
Lipid droplet functions beyond energy storage
Biochimica et Biophysica Acta (BBA) - 1862(10 Pt B),1260-1272
Michael A. Welte, Alex P. Gould (2017)
Drosophila Spidey/Kar Regulates Oenocyte Growth via PI3-Kinase Signaling
PLoS Genetics 12(8): e1006154
Einat Cinnamon, Rami Makki, Annick Sawala, Leah P. Wickenberg, Gary J. Blomquist, Claus Tittiger, Ze'ev Paroush, Alex P. Gould (2016)
Antioxidant Role for Lipid Droplets in a Stem Cell Niche of Drosophila
Cell 163, 340-353
Bailey AP , Koster G, Guillermier C, Hirst EMA, MacRae JI, Lechene CP, Postle AD, Gould AP (2015)
The development and functions of oenocytes
Annu Rev Entomol 59, 405-25
Makki R, Cinnamon E and Gould AP (2014)
Specialized hepatocyte-like cells regulate Drosophila lipid metabolism
Nature 445, 275-280
Gutierrez, E., Wiggins, D., Fielding, B. and Gould, A.P. (2007)
Developmental adaptations to malnutrition
|
Developing humans and other animals survive moderate nutrient deprivation by decreasing their overall growth rate and body size. During this adaptation, the growth of certain critical organs, such as the brain, is protected or spared at the expense of others. For example, in humans, fetal nutrient restriction can result in asymmetric intrauterine growth restriction (IUGR), characterised by undersized neonates with relatively large brains. Although brain sparing has been well documented in mammals for over 50 years, the underlying molecular mechanisms remain unclear.
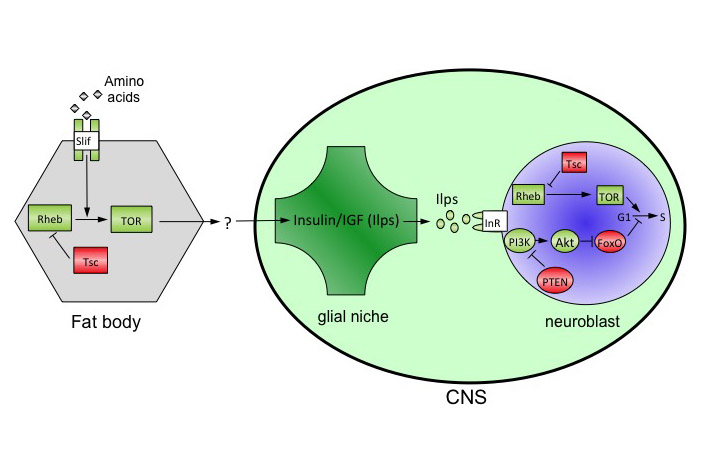
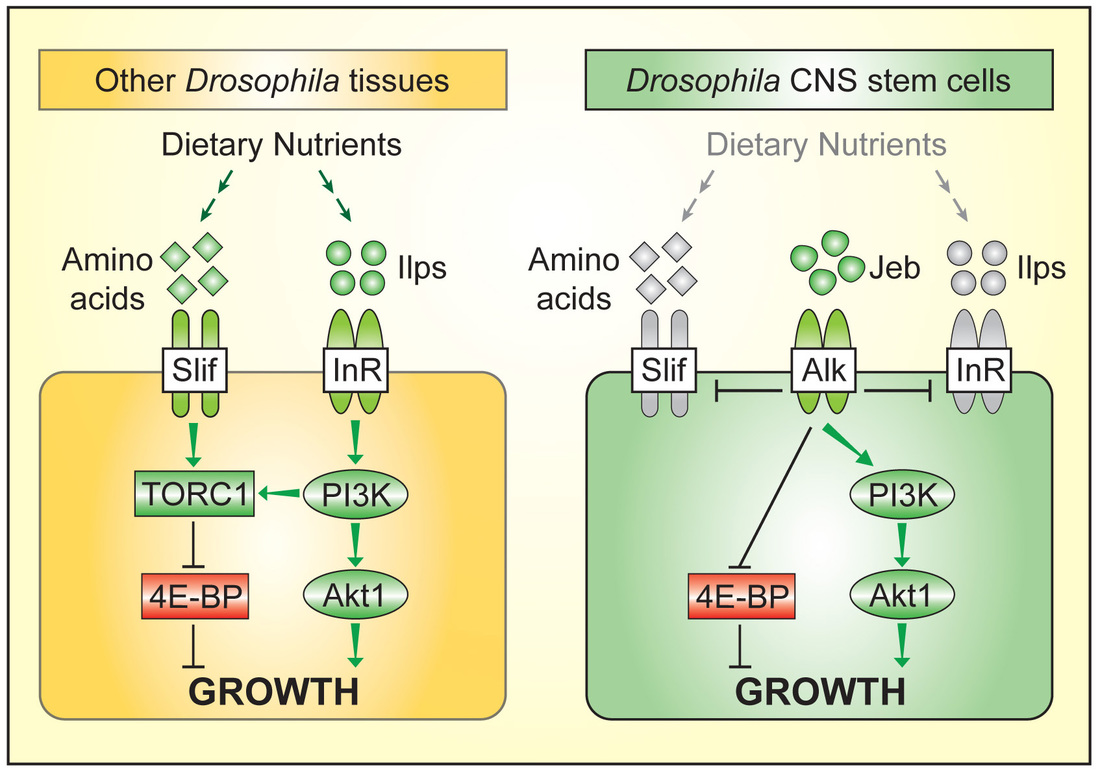
Work in our lab showed that brain sparing is present in Drosophila. We used the power of clonal analysis and other genetic techniques in Drosophila to show that CNS growth sparing during nutrient restriction involves the rewiring of nutrient sensing and TOR/PI3K signalling in neural stem cell-like progenitors called neuroblasts. This rewiring process is, however, only implemented at later stages of development. We found that it requires the activity of a specific receptor tyrosine kinase, Anaplastic lymphoma kinase (Alk). Alk stimulates PI3K signalling in neuroblasts, regardless of whether dietary nutrients are present or absent. This is because glial cells of the neuroblast niche constitutively secrete the ligand for Alk, Jelly belly. Neuroblast divisions are thus freed from nutrient-dependent growth components such as Insulin-like peptides. We are currently searching for additional growth sparing mechanisms in the CNS and in other tissues. In humans, Alk is an oncogene that is frequently hyperactivated in human large cell lymphomas, non-small cell lung cancers, neuroblastomas and other tumours. Alk is therefore a molecular link between the genetic networks driving selective tissue growth in brain sparing and in some cancers. We have established a mouse model for brain sparing at The Crick and are currently testing which brain sparing genes are functionally conserved between insects and mammals. |
Alk kinase domain bound by a small molecule inhibitor
|
Relevant publications:
Developmental diet regulates Drosophila lifespan via lipid autotoxins
Nature Communications 8, Article number: 1384
M. Irina Stefana, Paul C. Driscoll, Fumiaki Obata, Ana Raquel Pengelly, Clare L. Newell, James I. MacRae, Alex P. Gould (2017)
Protection of Neuronal Diversity at the Expense of Neuronal Numbers during Nutrient Restriction in the Drosophila Visual System
Cell Reports 3, 587-94
Lanet, E, Gould, AP and Maurange C. (2013)
Anaplastic Lymphoma Kinase Spares Organ Growth during Nutrient Restriction in Drosophila
Cell 146, 435-447
Cheng, L., Bailey, A., Leevers, S., Ragan, T., Driscoll, P. and Gould, A.P. (2011)
Fat cells reactivate quiescent neuroblasts via TOR and glial insulin relays in Drosophila
Nature 471, 508-512
Sousa-Nunes, R, Yee, LL and Gould, A.P. (2011)
Developmental diet regulates Drosophila lifespan via lipid autotoxins
Nature Communications 8, Article number: 1384
M. Irina Stefana, Paul C. Driscoll, Fumiaki Obata, Ana Raquel Pengelly, Clare L. Newell, James I. MacRae, Alex P. Gould (2017)
Protection of Neuronal Diversity at the Expense of Neuronal Numbers during Nutrient Restriction in the Drosophila Visual System
Cell Reports 3, 587-94
Lanet, E, Gould, AP and Maurange C. (2013)
Anaplastic Lymphoma Kinase Spares Organ Growth during Nutrient Restriction in Drosophila
Cell 146, 435-447
Cheng, L., Bailey, A., Leevers, S., Ragan, T., Driscoll, P. and Gould, A.P. (2011)
Fat cells reactivate quiescent neuroblasts via TOR and glial insulin relays in Drosophila
Nature 471, 508-512
Sousa-Nunes, R, Yee, LL and Gould, A.P. (2011)
New techniques for metabolic research
|
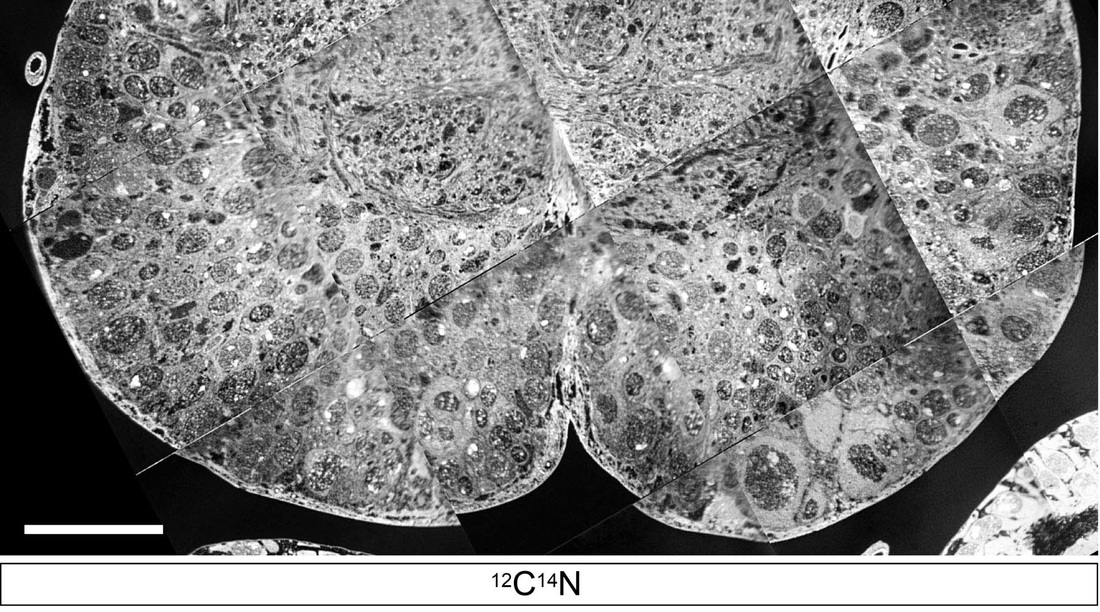
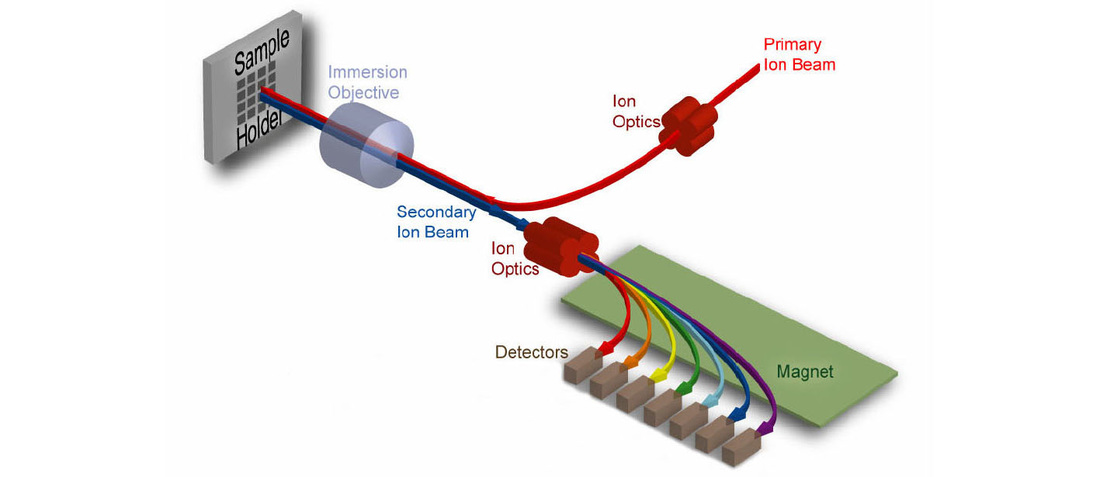
Methods for the analysis of metabolites by NMR and mass spectrometry are now becoming well established but technical limitations still remain. For example, our understanding of metabolism has been greatly hampered by an inability to image and to measure metabolites in subcellular compartments or even in individual cell-types within complex tissues. We have therefore developed, in collaboration with Claude Lechene (Harvard Medical School), new imaging mass spectrometry methods for the quantitation of metabolism with 50 nm spatial resolution. We anticipate many future applications for this powerful new technology and, thus far, have used it to measure the incorporation of dietary and newly synthesized fatty acids into individual lipid droplets of CNS, fat body and midgut cells.
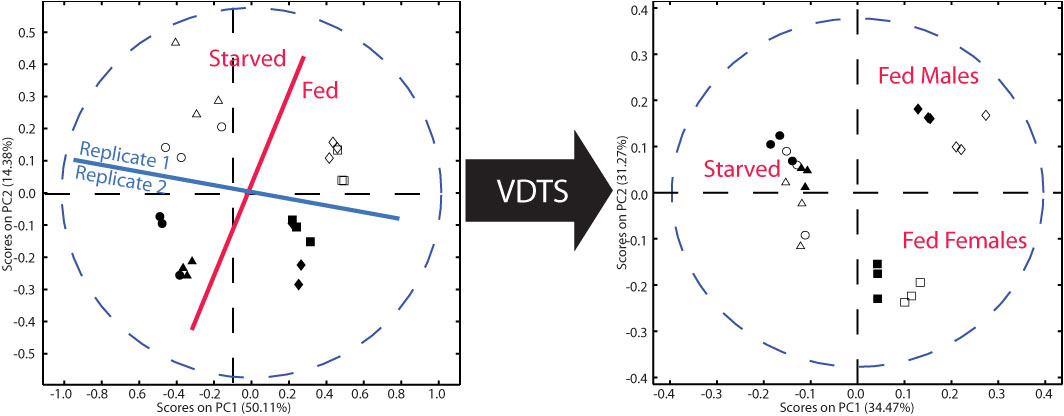
A serious limitation in metabolomics is the difficulty of measuring concentrations of metabolites in sub-microlitre quantities of fluids where the sample volume is not accurately known. To overcome this issue, we developed a technique called volume determination with two standards (VDTS). This approach permits the back calculation of absolute metabolite concentrations from small biological samples of unknown volume. As a proof-of-principle, we applied VDTS to an NMR analysis of polar metabolites from the hemolymph (blood) of tiny Drosophila larvae. In biological contexts where total signal strength differs considerably between samples, we found that VDTS greatly outperforms the widely used analytical method of probabilistic quotient normalization (PQN). We are greatly assisted in developing new mass spectrometry and NMR methods by James MacRae and Paul Driscoll of the Metabolomics Science and Technology Platform at The Crick. |
Relevant publications:
Sub-cellular Imaging of Metabolites with 3D OrbiSIMS
Cell Systems 5(6), 534-537.
Ian Gilmore, Andy West, Morgan Alexander, Alex Gould (2017)
Stable isotope analysis of dynamic lipidomics
Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids 1862 (8), 792-796
Joost Brandsma, Andrew P. Bailey, Grielof Koster, Alex P. Gould, Anthony D. Postle (2017)
Antioxidant Role for Lipid Droplets in a Stem Cell Niche of Drosophila
Cell 163, 340-353
Bailey AP , Koster G, Guillermier C, Hirst EMA, MacRae JI, Lechene CP, Postle AD, Gould AP (2015)
Volume determination with two standards allows absolute quantification and improved chemometric analysis of metabolites by NMR from submicroliter samples
Anal. Chem. 85, 12046-54
Ragan TJ, Bailey AP, Gould AP and Driscoll PC (2013)
Multi-isotope imaging mass spectrometry quantifies stem cell division and metabolism
Nature 481, 516-519
Steinhauser, M.L, Bailey, A.P, Senyo, S.E, Guillermier, C, Perlstein, T.S, Gould, A.P, Lee, R.T and Lechene, C.P. (2012)
Relevant publications:
Sub-cellular Imaging of Metabolites with 3D OrbiSIMS
Cell Systems 5(6), 534-537.
Ian Gilmore, Andy West, Morgan Alexander, Alex Gould (2017)
Stable isotope analysis of dynamic lipidomics
Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids 1862 (8), 792-796
Joost Brandsma, Andrew P. Bailey, Grielof Koster, Alex P. Gould, Anthony D. Postle (2017)
Antioxidant Role for Lipid Droplets in a Stem Cell Niche of Drosophila
Cell 163, 340-353
Bailey AP , Koster G, Guillermier C, Hirst EMA, MacRae JI, Lechene CP, Postle AD, Gould AP (2015)
Volume determination with two standards allows absolute quantification and improved chemometric analysis of metabolites by NMR from submicroliter samples
Anal. Chem. 85, 12046-54
Ragan TJ, Bailey AP, Gould AP and Driscoll PC (2013)
Multi-isotope imaging mass spectrometry quantifies stem cell division and metabolism
Nature 481, 516-519
Steinhauser, M.L, Bailey, A.P, Senyo, S.E, Guillermier, C, Perlstein, T.S, Gould, A.P, Lee, R.T and Lechene, C.P. (2012)